For much of our existence, pathogenic bacteria have had the upper hand on humanity. To put things into perspective, the average life expectancy in early 1900’s United States was a mere 47 years old, with the number one and number two causes of death being pneumonia and tuberculosis. Then, an accident changed everything. Alexander Fleming left some petri dishes laying around that had the bacterium Staphylococcus aureus growing on it, which got contaminated by a mold that could kill the bacteria. Further investigation revealed a microscopic war going on, with the fungal cell “shooting” the bacterial cell with a small compound called penicillin. It didn’t take us long for us humans to utilize this little “magic bullet” to our own advantage by mass-producing penicillin as an antibiotic. The discovery of penicillin catapulted antibiotic discovery, resulting in a “Golden Age” of antibiotic development. Not coincidentally, life expectancy increased about 30 years on average, and today our primary causes of death in the U.S. are heart disease followed by cancer. Thanks to antibiotics we overcame one of humanity’s biggest foes: bacterial pathogens. Or so we thought…
The “Golden Age” didn’t last long. By treating any ailment with antibiotics, we allowed for the survival of bacteria that possessed some resistance to the antibiotic. Maybe they could spit the drug back out, or the bacteria could shield themselves from the antibiotic. Whatever the resistance mechanism may be, the number of resistant bacterial pathogens is on the rise. Disconcertingly, the number of new antibiotics coming out of the drug discovery pipeline is steadily decreasing (Figure 1).
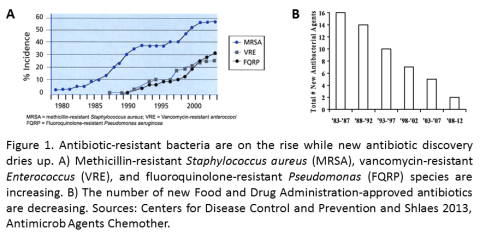
As you can imagine, superbugs increasing + antibiotic arsenal decreasing = recipe for disaster. We are now at risk of returning to the pre-antibiotic era and the diseases and epidemics of that time.
So what do we do? We’ve got to get creative. If you wanted to stop a zombie from chasing you, you probably wouldn’t kick it in the shin, right? You’ve got to go for the essential parts—the brain, the heart, or at least get rid of its legs so it can’t run after you. Within the bacterial cell, there are equivalents to these essential zombie parts, called essential pathways. If we can block these essential pathways within the bacterial cell using small molecule inhibitors, we have a chance to kill or at least slow down the bad bugs. Current antibiotics target only a handful of known essential pathways, yet there are dozens that remain unexploited; one such unexploited essential bacterial pathway is the process of RNA turnover (Figure 2).
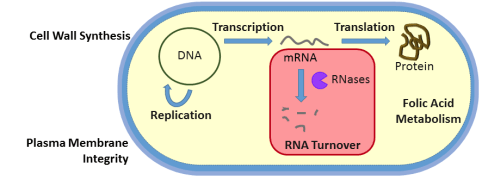
Within the bacterial cell, there is a general assembly line that starts by reading the DNA blueprint. Then the RNA middleman delivers the message, which is made into the functional protein—usually the enzymes that do a lot of the work in the cell. Once the RNA middleman has done its job, it has to be broken down and recycled for new RNA synthesis. The RNA turnover pathway is required for all life forms to live, but bacteria degrade their RNA quite a bit differently than humans do. Because of the differences in RNA turnover, we could specifically target only the bacterial enzymes that carry out these processes without harming the infected human.
And this is where I begin my story. My graduate studies were carried out under the mentorship of Paul Dunman, Ph.D. Our laboratory performed a small molecule screen for inhibitors to the essential S. aureus ribonuclease (RNase) RnpA. Within Gram-positive bacterial cells, RnpA is involved in two different processes; it acts as a member of the degradosome, to help with RNA turnover. RnpA is also a member of the ribonucleoprotein (RNA+protein team) complex RNase P, which processes transfer RNAs (tRNAs) involved in protein synthesis (Figure 3). My project was to further characterize and improve two promising S. aureus RnpA inhibitors, RNPA1000 and RNPA2000.
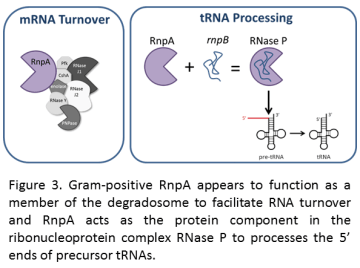
Before I get into the results of my graduate work, let me say that hindsight is 20/20. In retrospect, I can describe my research as a logical progression of A leads to B leads to C and so on, making it seem like I just tip-toed through the tulips to get useful data. In reality, A leads to F, leads to Z, leads to banana, and then there’s a dead end. In short, I struggled in graduate school. A lot. Three or four unsuccessful projects passed through my hands, resulting in several years of basically unpublishable work. It was frustrating and challenging, but my boss Paul kept (lovingly) kicking my butt to encourage me to get back up and try again. The last year and a half or so of my graduate studies is when something crazy happened—my experiments were working! I showed that both RNPA1000 and RNPA2000 could block S. aureus RNA turnover. While RNPA1000 was bacteriostatic (limited the growth of S. aureus), RNPA2000 was bactericidal (killed S. aureus). Further investigation demonstrated that only RNPA2000 could inhibit tRNA processing within S. aureus, providing evidence it could block both activities of RnpA (as a member of the degradosome and as a member of RNase P), likely contributing to its bactericidal effects. Importantly, RNPA2000 was not toxic to human cells, illustrating that it had potential to be used in people. Read more in the full manuscript.
These experiments illustrate that RnpA inhibitors have the potential to be antimicrobial agents, representing a potential new class of antibiotics. Importantly, there is hope that the drug discovery pipeline has not dried up. Yes, the low-lying fruit on the antibiotic tree has been picked, but there are lots of other goodies up in that tree, and it’s our job as scientists to climb up there and get ‘em!